Without well-functioning antibiotics many medical procedures, such as surgery or transplantation would be too risky, and thus impossible to perform. Common infections, today considered easily treatable, might become deadly.
Antibiotics hit bacteria where it hurts, by inhibiting key cellular processes. More than half of all antibiotics used today target the essential cellular process of protein production. All cellular proteins are made by a molecular machine, the ribosome, which is therefore the single most important antibiotic target.
Bacteria are constantly developing ingenious and varying ways to avoid the effect of the antibiotic. The important mechanisms that are targeted by the antibiotic can change so that the antibiotic no longer has an effect. Once this resistance has evolved, because it’s encoded in DNA, bacteria can pass it on to their descendants or cousins.
A picture is worth a thousand words
In Sweden antibiotic resistance is a problem, though not as serious as it can be in other parts of the world. And the problem is growing. To develop new antibiotics that will be able to overcome resistance, it is essential to understand how the resistance works.
– A crucial step in understanding is through visualisation of the mechanism on the molecular level with protein structures. As they say, a picture is worth a thousand words, says Associate Professor Vasili Hauryliuk who has led the study, in collaboration with Professor Daniel N Wilson and his team at University of Hamburg.
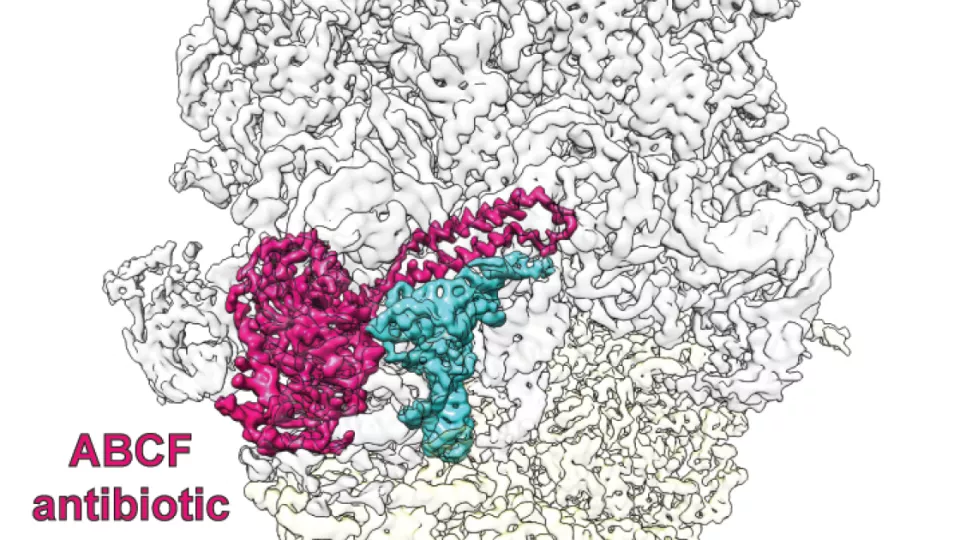
The researchers uncovered the molecular details of a widespread antibiotic resistance mechanism mediated by so-called ABCF ATPases. These resistance factors protect the ribosome from several classes of clinically important antibiotics. Using cryogenic electron microscopy (cryo-EM) the teams have generated structures of ABCF resistance factors from several important pathogens; Listeria monocytogenes, Staphylococcus aureus and Enterococcus faecalis, bound to the ribosomes they protect.
– Understanding these structures is the first step towards developing new antibiotics that are harder to break down, and can overcome ABCF-mediated resistance. When you see how the defence mechanism works, you can try to modify the existing antibiotic so it can outsmart the resistance factor, and the bacteria, says Associate Profesor Gemma C. Atkinson, a contributor to the project.
In their study the researchers used a powerful structural biology approach, cryo-EM (single-particle cryogenic electron microscopy).
– This is a very important technique that evolves rapidly and allows you to see molecules in exceptional detail. Cryo-EM is currently superseding the default structural approach: X-ray crystallography, because it does not require crystallisation of your sample, the sample is imaged directly. Very small amounts are needed and one can work with a mixture of different molecules, explains Gemma C. Atkinson
All of the data were collected at the national Swedish cryo-EM facility in Umeå. Right now the facility has two nodes, one in Umeå and one in Stockholm. Uppsala University is purchasing a cryo-EM instrument.
– Unfortunately, right now there is no cryo-EM instrument in Lund, but we hope this will soon change. The Department has requested funding from the faculty for purchase of cryo-EM.
It is crucial for Lund’s competitiveness in structural biology, both nationally and internationally, says Vasili Hauryliuk.